Operation and assembly of the bacterial flagellum.
Chapter three describes the three main parts of a flagellum.
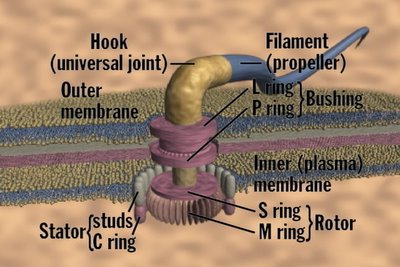
Basal body, hook and filament.
The filament is hollow and made of flagellin, and rigid.The flagellin subunits assemble spontaneously to give the filament.
The hook is at the base of the filament and appears to act like a "universal joint" when the flagella is rotating.
The basal body is imbedded in the cell membranes, and include some 15 different proteins.
It acts as an electric rotating motor driven by PMF.
The flagellum is assembled in a stepwise fashion. First the basal body is inserted into the membrane, next the hook, finally the filament.
Units of flagellin reach the tip through its hollow centre.
Regulatory Loops.
In E. coli synthesis of flagellin subunits is coupled to their assembly at the tip by a special inhibitor of flagellin synthesis. When the basal body is inserted into the membrane it accomplishes secretion of this inhibitor into the growth medium which thus allows filament subunit synthesis to start.
Proteins needed to allow the flagella to rotate are added late in organelle assembly.
Study Questions:
What is PMF and where does it come from?
How were the stages of flagellum assembly first identified?

2 Comments:
Proton motive force is a term analogous to electromotive force, which refers to the difference in voltage which causes an electric current to flow. (The term force is misleading because the unit of emf is the volt).
Proton motive force is an electrochemical potential created by a proton gradient across a lipid membrane, creating a charge difference between sides. This difference is also measured in volts (transmembrane potential)
H+ from the side with greater [H+] moves across the membrane to that of lower [H+], in an exergonic process which releases energy.
This electrochemical energy is captured by ATP synthase, and transformed into mechanical energy, rotating subunits which assemble ATP molecules.
Proton gradients are also made by bacteria by running ATP synthase in reverse, used to drive flagella.
Jeremy R. 9
Good comments. PMF can have both a pH difference and an electical charge (voltage) difference component in various proportions. It is the difference in chemical potential of protons. Thus Proton-motive is a useful word to refer to it.
Insides of cells are often more alkaline than the exterior, at low exterior pH.
Post a Comment
<< Home