This discussion thread is to assist people who want to write an assignment essay about molecules secreted by
Pseudomonas aeruginosa, a gram negative bacterium commonly found in natural waters, that can also be a highly antibiotic resistant human pathogen.
The same general comments made in the previous thread about how to start researching the topic of cell division apply also here. Use NCBI-PubMed, and put your comments, questions, and discoveries in the comments please.
This current posting is designed to explain a different way of quickly finding the scientific status of a topic. I suggest here to try exploiting the
Introduction section of a relevant and very recent research paper.
These Introduction sections are
actually short up to date reviews by very competant scientists. They may be concise, but they are packed with the benefits of special expertise, and can be much more up to date than the most recently available mini-review, or recent longer review article.
The tricky challenge is to quickly locate a suitable recent paper.
You need to start somewhere close to the topic and leverage off that starting point.
Microbe Pundit has already suggested that
multi-drug efflux pumps of Pseudomonas are involved in secretion of many different compounds. Why not start at this factoid?
He's mentioned already that Greenberg's journalistic style article
Pumping up the versatility in a Nature journal is a good place to start discovering what's been learned about Pseudomonas.
But when reading Greenberg's article, you will find there is surprizingly little detail about molecule secretion, and it's a few years out of date.
But it does have the words :
RND pump multidrug effluxmentioned so plug them into an NCBI-Pub Med search (Google NCBI first if you're lost for a URL).
With this literature search short-cut you come quickly (if you use a bit of intelligence) to:
Nehme D, Li XZ, Elliot R, Poole K.
Related Articles, Links
Free in PMC Assembly of the MexAB-OprM
multidrug efflux system of
Pseudomonas aeruginosa: identification and characterization of mutations in mexA compromising MexA multimerization and interaction with MexB.
J Bacteriol. 2004 May;186(10):2973-83.
The bolded words show why Pundit thought this was gold.
Now get to the full text version of the J Bacteriol paper via your friendly university library or better, from
FREE IN PMC over the internet anywhere with a terminal. Remember to have your USB thumb drive with you at the computer. (An USB MP3 player will also suffice, you just have to delete all those Kylie Minogue MP3s.)
With this you get to:Pseudomonas aeruginosa is an opportunistic human pathogen characterized by an innate resistance to multiple antimicrobials (21). The resistance has historically been attributed to the presence in this organism of an outer membrane (OM) of low permeability (55), but it is increasingly clear that resistance owes much to the operation of broadly specific, so-called multidrug efflux systems (58-60, 64) that work synergistically with limited OM permeability (18, 40, 60). Several multidrug efflux systems in
P. aeruginosa have been described to date (61), although the major system contributing to intrinsic multidrug resistance is encoded by the mexAB-oprM operon (38). Hyperexpression of this system also occurs in so-called nalB (27, 28, 75, 88)- and nalC (75, 88)-type multidrug-resistant mutants. MexAB-OprM accommodates a broad range of structurally diverse antimicrobials, including dyes, detergents, inhibitors of fatty acid biosynthesis, organic solvents, disinfectants, and clinically relevant antibiotics (10, 34, 37, 39, 41, 42, 48, 70, 74, 74, 76),
and is implicated in the export of homoserine lactones involved in quorum sensing (17, 57) and, possibly, virulence factors (22).Evans, K., L. Passador, R. Srikumar, E. Tsang, J. Nezezon, and K. Poole. 1998.
Influence of the MexAB-OprM multidrug efflux system on quorum-sensing in Pseudomonas aeruginosa. J. Bacteriol. 180:5443-5447.
Pearson, J. P., C. Van Delden, and B. H. Iglewski. 1999.
Active efflux and diffusion are involved in transport of Pseudomonas aeruginosa cell-to-cell signals. J. Bacteriol. 181:1203-1210.
Hirakata, Y., R. Srikumar, K. Poole, N. Gotoh, T. Suematsu, S. Kohno, S. Kamihira, R. E. Hancock, and D. P. Speert. 2002.
Multidrug efflux systems play an important role in the invasiveness of Pseudomonas aeruginosa. J Exp. Med. 196:109-118.
The MexAB-OprM efflux system, like the other tripartite Mex efflux systems in
P. aeruginosa, consists of an inner membrane (IM) drug-proton antiporter of the resistance-nodulation-cell division (RND) family (MexB), an OM channel-forming component (OprM; also called OM factor [OMF]), and a periplasmic membrane fusion protein (MFP) (MexA) (58, 86). Crystal structures have not yet been reported for any of the efflux components of P. aeruginosa,
although structures are available for the homologous OM (TolC [35]) and RND (AcrB [52]) components of the Mex-like AcrAB-TolC multidrug efflux system of Escherichia coli. The TolC channel is a trimer and spans both the OM (as a ß-barrel) and periplasm (as a {alpha}-helical barrel) (35). Measuring 140 Å in length, the channel is open at the distal (extracellular) end and tapers almost to a close at the proximal (periplasmic) end, which likely interacts with the RND component, AcrB (35). Modeling studies suggest that OprM adopts a similar structure (43, 80). The AcrB RND component also exists as a trimer, composed of a 50-Å-thick transmembrane region and a 70-Å headpiece that protrudes into the periplasm (52). This headpiece has a funnel-like opening at the top that is connected to a central cavity at the bottom, which, in turn, opens to the periplasm via three vestibules that likely play a role in substrate recognition by and/or access to the pump (52). Indeed, several studies highlight the role played by the periplasmic portion of the RND transporters of
E. coli and
P. aeruginosa in substrate (i.e., drug) recognition (14, 15, 45, 53, 79, 83). A high-resolution crystal structure for an MFP efflux component is not yet available, although preliminary studies indicate that, e.g., AcrA is also likely trimeric (85) and that monomer AcrA is a highly asymmetric, elongated molecule of sufficient length to span the periplasm (4, 84).
Koronakis, V., A. Sharff, E. Koronakis, B. Luisi, and C. Hughes. 2000.
Crystal structure of the bacterial membrane protein TolC central to multidrug efflux and protein export. Nature 405:914-919.
Bold indicates the words that provide leads to interesting aspects of secretion by Pseudomonas.
A mini-review essay topic could well focus entirely on homoserine lactone or maybe protein virulence factor secretion by Pseudomonas, and could involve careful reading, and re-reading, of 2 to 3 relevant review papers and 4 to 6 good research papers on the topic. That's enough work for an assigment.
PS Remember to
ITALICISE all those species binomials in your final assignment. Cite references professionally too.
And finally, trust your luck .
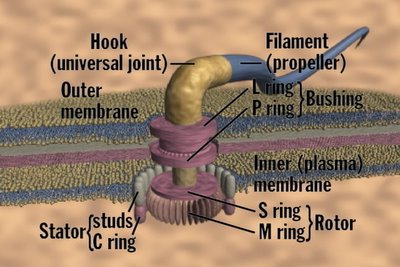
In an effort to brighten up this post with a colourful image (posted at the top), I did a google image search for bacterial multi-drug efflux pumps and found
this.