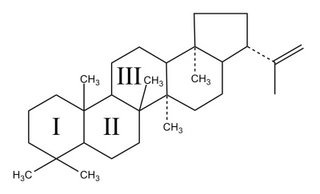
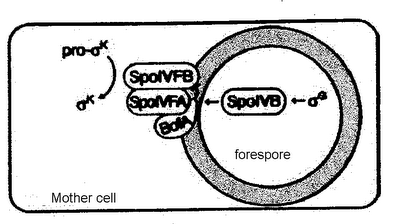
Where the wild-things are, and where avermectin comes from.
Geek territory?
Many of us would prefer to just read the paper's abstract.
Thus for those who want a kindler gentler commentary on the metabolic capabilities of Streptomyces, Fam has kindly provided the following. Thanks Fam.
Streptomyces avermitilis
Fam W. N
Abstract
Streptomyces avermitilis is the producer of Avermectin, an anthelmintic macrolide which was isolated by Omura et al. Having the largest bacterial genome sequence, S.avermitilis has an interesting ability to produce a variety of secondary metabolites. Genome analysis has led to identification of total of 30 gene clusters involved in secondary metabolite biosynthesis in S.avermitilis, and more than half of them are located near at its chromosome ends. In this review, structure, general characteristics and practical applications of Avermectin will be addressed as well as analysis of these gene clusters in S.avermitilis which give insight into complex diversity of secondary metabolites and their physiological roles.1. Introduction
Streptomyces is a genus of gram-positive Actinomycete bacteria. These filamentous bacteria are found mainly in soil and marine habitats (Lamb et al., 2003). The unique characteristic of streptomycete bacteria is that they are able to exhibit a complicated life cycle involving formation of a lawn of aerial hyphae on the colony surface that stands up into the air and differentiates into chains of spore (Kelemen and Buttner, 1998). This process is highly controled and requires specialized coordination of metabolism. Because of diversity of their secondary metabolic pathway, streptomycetes are being extensively studied for their biologically active products which are widely used in human and veterinary medicine, as antibiotics, antiparasitic agents, as well as other pharmaceutical activities (Ikeda et al., 2003).
Streptomyces avermilitis is one soil bacterium in this genus. Streptomyces avermilitis was first isolated by Omura et al. of the Kitasato Institute from the soil sample which collected in Ito city, Shizuoka Prefecture, Japan (Burg, 1979). It has a large genome (9.02 Mb) and its chromosome exists as linear which is similar in the cases of other bacteria of the same genus. Taxonomic studies revealed its morphology with sporophores forming spirals as side branches on aerial mycelia (Fig. 1). The spore surface was smooth by observation made under electron microscopy (Burg, 1979). Merck Sharp and Dohme discovered the effective activity of Avermectin in a broth of S.avermilitis by chance. Testing found that it was non-toxic towards the lab mice. It also showed potentially as an anti-helminthic agent, and the activity was ten-fold higher than any synthetic antihelminthic agent (Demain, 1983).
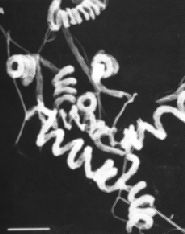 Fig. 1: (Left) Scanning electron micrograph of Streptomyces avermitilis, from http://www.nih.go.jp/saj/atlas/sample.html. (Right) Transmission electron micrograph showing its smooth surface (Burg, 1979) [not shown].
Fig. 1: (Left) Scanning electron micrograph of Streptomyces avermitilis, from http://www.nih.go.jp/saj/atlas/sample.html. (Right) Transmission electron micrograph showing its smooth surface (Burg, 1979) [not shown].Avermectin is well-known as an excellent anthelminthic agent and it is highly against a broad spectrum of nematode and arthropod parasites. Due to its superior activity, avermectin market exploded during 1980s and reached U.S $1 billion at the end of 1990 (Hwang et al., 2003). Its semisynthetic derivative, Ivermectin is also widely used in veterinary and agricultural fields (Ikeda et al., 1999).
Genome sequence completion has revealed 30 gene cluster in Streptomyces avermitilis that are proposed to be involved in biosynthesis of secondary metabolites (Ikeda et al., 1999). This indicates that there could be a large number of metabolic pathways and hence a variety of potential bioactive compounds that could be as useful as Avermectin awaiting discovery.
2. Avermectin
Avermectins are a complex of macrocyclic lactone derivatives which in contrast to macrolide or polyene antibiotics, lacking significant antibacterial or antifungal activity (Burg, 1999).
2.1 Biosynthesis of Avermectin
Basically, Avermectin biosynthesis can be divided into 3 stages. The first stage is the formation of polyketide-derived initial aglycon. The avermectin polyketide synthase, PKS uses a range of acyl units (Ikeda et al., 1999). The starting acyl group for synthesis of Avermectin derived from valine to form a components and isoleucine to form b components (Lamb et al., 2003). The second stage involves the modification of initial aglycon to generate Avermectin aglycons. This is formed by extension of started unit with addtition of 7 acetate and 5 propionate units and involves a few modification steps which include ketoreduction, methylation and furan ring closure. The final stage involves O-glycosylation at C13 and C4’ to form Avermectins from Avermectin aglycons. This glycosylation step is performed by deoxythymidine diphosphate (dTDP)-oleandrose.
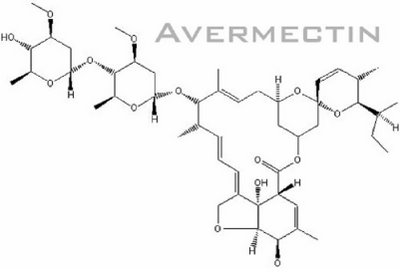
(figure from http://www.plantaanalytica.com/avermectins.htm)
2.2 Mode of action of Avermectin
Avermectin and its chemically derived compound, Ivermectin can affect a variety of ligand- and voltage-gated chloride channels (Bloomquist, 1993). It has high affinity to bind glutamate-gated chloride ion channels which is found in peripheral nervous system of invertebrates, establishing its selective activity against human parasites. Binding of Avermectin in this ion channel block electrical activity in vertebrate and invertebrate nerve and muscle cells by increasing the permeability of cell membrane to chloride ions. A consequence of this is the hyperpolarization followed by paralysis and death of the parasite.(Santoro et al., 2003) In normal condition, Avermectin and Ivermectin does not cross mammalian blood-brain barrer and human are spared from serious central nervous system effects that brought by this drug.
2.3 Practical application of Avermectin and Ivermectin
Initially, Avermectin was used as insecticide to protect the crops. It was later used to treat infection in livestock and domestic animals caused by nematodes and arthropods (Santoro et al., 2003). Most importantly, Avermectin has been made as promising drug in human onchocerciasis. It has conferred some protection to a huge population in sub-Saharan Africa in which the disease called river blindness was prevalent at a certain point of time (Hopwood, 2003).
It has become evident that Ivermectin is effective in treating other filariases such as loiasis, bancroftian filariasis and other intestinal nematodes. Inoculation of this drug successfully inhibits the maturation of larvae at certain developmental stages of filarial species. This could be used to protect the domestic animals from diseases such as heartworm disease (Campbell et al., 1983). Some nematodes can enter hypobiosis. However, this does not deter the action of Ivermectin unlike most of other anthelmintic agents. Thus, it is considered strong weapon in the field of parastic dermatology, where it can act against its target at any stage of development (Pascal et al., 2003).
Ivermectin also emerges as an oral antiscabietic to treat scabies. It is shown to be as safe and effective as topical antiscabietics. Recent report revealed that all groups of population tested show good response to Ivermectin in the treatment scabies. These populations include immunocompromised, immunocompetent as well as in other high-risk populations such as those with Down’s syndrome (Santoro et al., 2003).
3. Review of potential of Streptomyces avermitilis as producer of a wide variety of secondary metabolites .
Soil contains highly diversity of bacterial communities. It is a complex environment in which bacteria will encounter chemical, physical as well as biological stress. To combat the stress and compete with other microorganisms, Streptomyces avermilitis which is non-motile possess a large number of genes encoding nutritional enzymes, transport proteins, cell regulators and other useful secondary metabolites (Challis and Hopwood, 2003).
Genome analysis revealed Streptomyces avermitilis has at least 30 kind of secondary metabolite gene clusters in chromosome (Ikeda et al., 2003). Twenty five clusters that have studied earlier involved in biosynthesis of compounds such as carotenoid, melanin, polyketide, siderophore and peptides (Ikeda et al., 1999). Recent studies identified 5 more gene clusters which involved in biosynthesis of terpene and polyketide compounds (Ikeda et al., 2003). The total length of these gene clusters was estimated to be 594 kb, which indicate that about 6.6% of S.avermilitis genome is occupied by genes involved in biosynthesis of secondary metabolites. Most of these gene clusters were observed to be located on the ends of the chromosome and contained many transposable elements. These transposases might be responsible for transferring some secondary metabolite genes into S.avermitilis via horizontal transfer and contribute to high production of secondary metabolites (Omura et al., 2001). Genetic studies suggest that its genome might have undergone evolution by acquisition of novel gene functions in order to survive in extremely variable soil environment with rapid changes in its physical conditions as well as to face tough competitors in obtaining available nutrients. (Ikeda et al., 2003).
Gene clusters analysis discovered that S.avermitilis has the ability to produce two anti-fungal compounds which are oligomycin and polyene macrolide. These two compounds are believed to potentially act synergistically against fungal competitor in its natural environment. (Challis and Hopwood, 2003). In addition, genes encode for two extracellular enzymes with glucanase and chitinase activities are found highly expressed in S.avermitilis. They also play important role against fungi. (Wu et al., 2005).
Cytochrome P450, CYP genes encode a family of heme-thiolate-containing enzymes. The CYP genes are normally located in macrolide antibiotic biosynthetic gene clusters. There are 33 cytochromes P450 (CYPs) found in newly completed S.avermitilis genome (Chater, 1989). It is predicted that 11 of this CYPs might be involved in biosynthesis of secondary metabolism and remaining CYPs might play a role in protecting S.avermitilis against toxic compound in soil environment (Lamb et al., 2003).
Siderophores are involved in transportation of iron in bacteria. Recent genome analysis also found a gene cluster in S.avermitilis which is presumably involved in biosynthesis of desferrioxamine derivatives (Ikeda et al., 2003). This can be used to treat acute iron poisoning especially in small children and proven to be an effective treatment for Ataxia-telangiectasia which is a genetic neurological disorder (Olivieri,1990).
The emergence of antibiotic resistant and new infectious diseases has brought a great demand in discovering novel antibiotics. Genome mining of S.avermitilis provides valuable information of this strain in biosynthesis of wide variety of secreted molecules/secondary metabolites which could be as useful as Avermectin and contributing to novel drug discovery. In addition, various bioinformatics resources such as BLAST (http://www.ncbi.nlm.nih.gov/blast/), ScanProsite ((http://ca.expasy.org/prosite/), enable us to do genome comparative analysis of streptomyces strains to gain insight the evolution of streptomyces family and conserved useful genes that contribute to novel antibiotics production which might advantageous not only to streptomyces itself but as well as in biomedical, pharmaceutical and biotechnological industry.
Acknowledgements
I thank Dr. M. Pundit for his valuable advice and guidance on different aspects of doing this proposal.
References
1. Bloomquist, J. R. 1993. Toxicology, mode of action, and target site-mediated resistance to insecticides acting on chloride channels. Mini Review, Comp. Biochem. Physiol 106: 301-314.
2. Burg, R.W. 1979. Avermectins, new family of potent antihelminthic agents: producing organism and fermentation. Antimicrob. Agents Chemother. 15:361-367.
3. Campbell, W.C., M.H. Fisher, E.O. Staphley, G. Albers-Schonberg, and T.A. Jacob. 1983. Ivermectin: A potent new antiparasitic agent. Sci. 221:823-828.
4. Challis, G. L., and D. A. Hopwood. 2003. Synergy and contingency as driving forces for the evolution of multiple secondary metabolite production by Streptomyces species. Proc. Natl. Acad. Sci. USA 100(Suppl. 2):14555-14561.
5. Chater, K.F. 1989. Multilevel regulation of Streptomyces differentiation. Trends Genet. 5:372–377.
6. Demain, A.L. 1983. New applications of microbial products. Sci. 219(4585):709-714.
7. Hopwood, D.A. 2003. The Streptomyces genome- be prepared! Nature Biotech. 21:505-506.
8. Hwang, Y.S., Kim, E.S., Biro S. and Choi C.Y. 2003. Cloning and Analysis of a DNA Fragment Stimulating Avermectin Production in Various Streptomyces avermitilis Strains. Appl. Envir. Microbiol. 69(2): 1263 - 1269.
9. Ikeda H, Ishikawa J, Hanamoto A, Shinose M, Kikuchi H, Shiba T, Sakaki Y, Hattori M, Omura S. 2003. Complete genome sequence and comparative analysis of the industrial microorganism Streptomyces avermitilis. Nat Biotechnol. 21;526-531.
10. Ikeda, H., T. Nonomiya, M. Usami, T. Ohta, and S. Omura. 1999. Organization of the biosynthetic gene cluster for the polyketide anthelmintic macrolide avermectin in Streptomyces avermitilis. Proc. Natl. Acad. Sci. USA 96:9509-9514
11. Kelemen, G.H., and Buttner, M.J. 1998. Initiation of aerial mycelium formation in Streptomyces. Curr. Opin. Microbiol. 1: 656–662.
12. Lamb, David C, Ikeda, Haruo, Nelson, David R, Ishikawa, Jun, Skaug, Tove, Jackson, Colin, Omura, Satoshi, Waterman, Michael R, Kelly, Steven L. 2003. Cytochrome p450 complement (CYPome) of the avermectin-producer Streptomyces avermitilis and comparison to that of Streptomyces coelicolor A3(2). Biochem Biophys Res Commun, 307(3):610-9.
13. Lamb, David C., Ikeda, Haruo, Nelson, David R., Ishikawa, Jun, Skaug, Tove, Jackson, Colin, Omura, Satoshi, Waterman, Michael R., Kelly, Steven L. 2003. Cytochrome p450 complement (CYPome) of the avermectin-producer Streptomyces avermitilis and comparison to that of Streptomyces coelicolor A3(2). Biochem Biophys Res Commu., 307(3):610-9.
14. Omura, S., Ikeda, H., Ishikawa, J., Hanamoto, A., Takahashi, C., Shinose, M., Takahashi, Y., Horikawa, H., Nakazawa, H., Osonoe, T., Kikuchi, H., Shiba, T., Sakaki, Y., and Hattori, M. 2001. Genome sequence of an industrial microorganism Streptomyces avermilitis: Deducing the ability of producing secondary metabolites. Proc. Natl. Acad. Sci. U. S. A. 98:12215–12220.
15. Pascal, D.G., Olivier C., and Caumes E. J. 2003. Ivermectin in dermatology. J. of Drugs in Derma. 2:327-313.
16. Santoro, A. F., M. A. Rezac, and J. B. Lee. 2003. Current Trend in Ivermectin Usage for Scabies. J. of Drugs in Derma. 2:397-401.
17. Wu, G., Culley, D. E. and Zhang, W. 2005. Predicted highly expressed genes in the genomes of Streptomyces coelicolor and Streptomyces avermitilis and the implications for their metabolism. Microbio. 15:2175–2187.
18. Olivieri NF, Koren G, Hermann C, Bentur Y, Chung D, Klein J, St Louis P, Freedman MH, McClelland RA, Templeton, DM. 1990. Comparison of oral iron chelator L1 and desferrioxamine in iron-loaded patients. Lancet 336: 8726
Labels: Biodiversity, Discovery